This page was produced as an assignment for Genetics 677, a undergraduate course at UW-Madison
Protein Phylogeny
________________________________________________________________________________________________________
To learn more about evolutionary conservation of CAPN3, phylograms were constructed with GeneBee and phylogeny.fr using multiple sequence alignments (MASs) generated by MUSCLE, ClustalW2, and T-Coffee. All trees and alignments were done using the default settings.
The initial MAS was found on HomoloGene (NCBI) and generated by MUSCLE* on 8 species (H sapiens, P troglodytes, B taurus, C lupus, M musculus, R norvegicus, D rerio, and G gallus), however several accessions had either been outdated, replaced or deleted. It was therefore necessary to search NCBI Protein for the most recently updated/stable accessions to use in phylogenetic analysis.
*The HomoloGene MUSCLE MAS can be found here
MUSCLE Alignment
ClustalW2 Alignment
Discussion
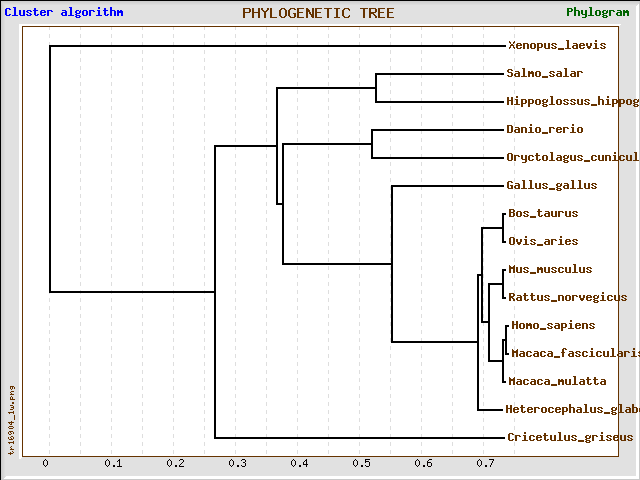
Multiple sequence alignments between CAPN3 protein homologs reveal that they are most conserved among primates (Macaca) and rodents (M musculus, R norvegicus, H glaber), with chinese hamster (C griseus) and rabbit (O cunelicus) less closely related, but more related than G gallus (according to the clustalw2 algorithm).
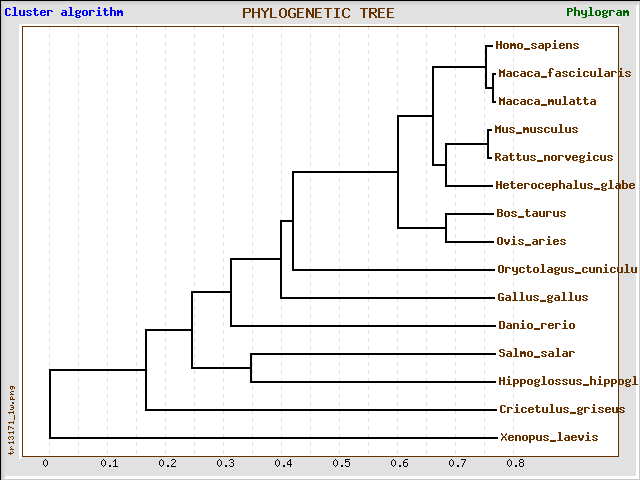
T-Coffee predicted H hippoglossus, S salar, and D rerio to have a much closer related capn3 to humans than the clustalw2 and MUSCLE alignments. All alignment software used seemed to show the same conservation among primates and rodents, however there were slight differences in clade members and phylogenetic distances among other species (especially D rerio and O cunelicus).
T-Coffee predicted H hippoglossus, S salar, and D rerio to have a much closer related capn3 to humans than the clustalw2 and MUSCLE alignments. All alignment software used seemed to show the same conservation among primates and rodents, however there were slight differences in clade members and phylogenetic distances among other species (especially D rerio and O cunelicus).
________________________________________________________________________________________________________
|
References
[1] Armougom F, Moretti S, Poirot O, Audic S, Dumas P, Schaeli B, Keduas V, Notredame C. Expresso: automatic incorporation of structural information in multiple sequence alignments using 3D-Coffee. Nucleic Acids Res. 2006 Jul 1;34(Web Server issue):W604-8. [2] Larkin MA, Blackshields G, Brown NP, Chenna R, McGettigan PA, McWilliam H, Valentin F, Wallace IM, Wilm A, Lopez R, Thompson JD, Gibson TJ and Higgins DG. ClustalW and ClustalX version 2 (2007) Bioinformatics 2007 23(21): 2947-2948. [3] Di Tommaso P, Moretti S, Xenarios I, Orobitg M, Montanyola A, Chang JM, Taly JF, Notredame C. T-Coffee: a web server for the multiple sequence alignment of protein and RNA sequences using structural information and homology extension. Nucleic Acids Res. 2011 Jul;39(Web Server issue):W13-7. Epub 2011 May 9. |
[4] TreeTop: Phylogenetic Tree Prediction (GeneBee)
[5] Edgar RC. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004 Mar 19;32(5):1792-7. Print 2004. |
This page was created by Pavlos Bousounis. Contact: [email protected]
Last Updated 2.23.2012
Last Updated 2.23.2012